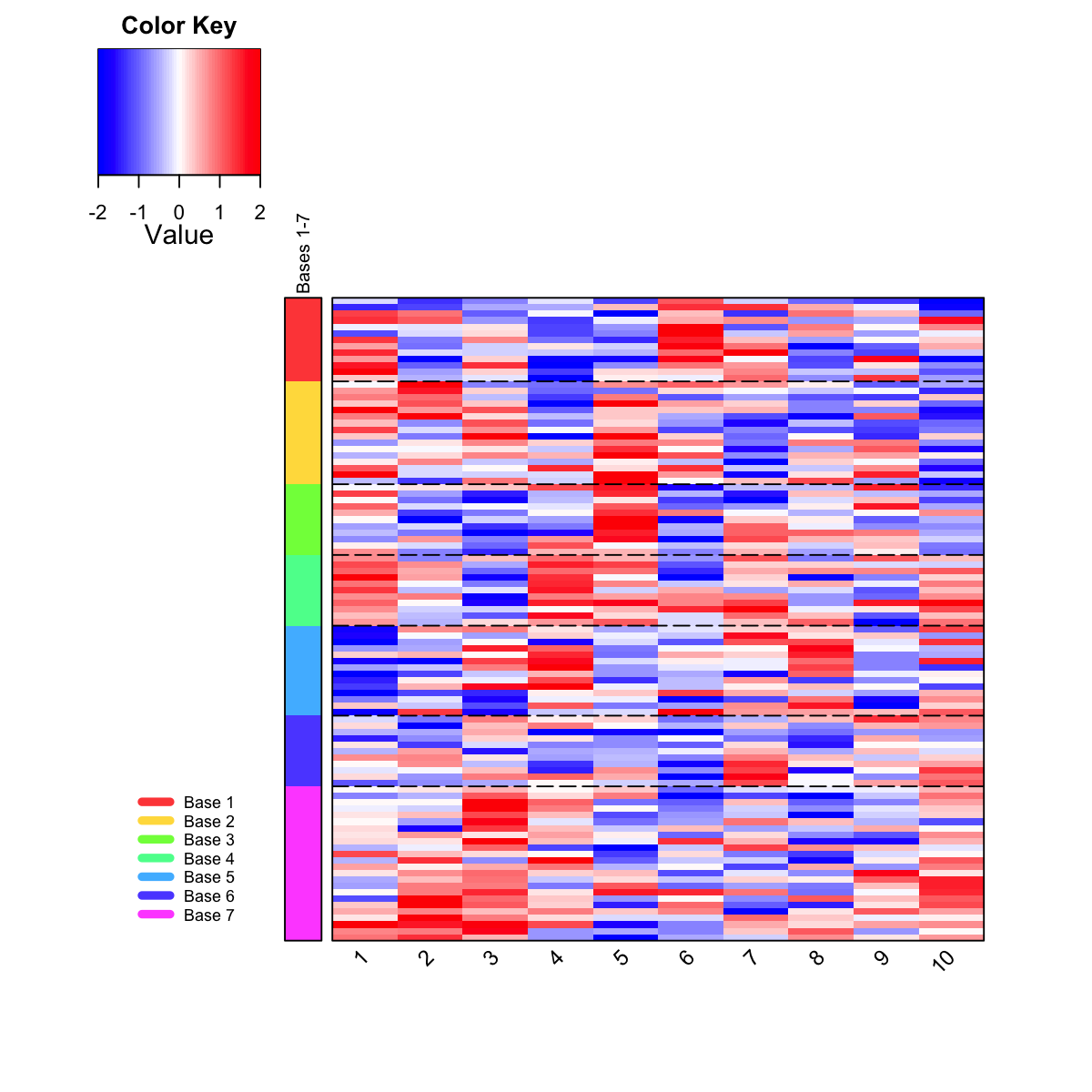
Function to visualise gene clusters/bases partitioned from a supra-hexagonal grid using heatmap
Description
visDmatHeatmap is supposed to visualise gene clusters/bases
partitioned from a supra-hexagonal grid using heatmap
Usage
visDmatHeatmap(sMap, data, sBase, base.color = "rainbow", base.separated.arg = NULL,
base.legend.location = c("none", "bottomleft", "bottomright", "bottom", "left",
"topleft", "top", "topright", "right", "center"), reorderRow = c("none",
"hclust", "svd"), keep.data = FALSE, ...)
Arguments
- sMap
- an object of class "sMap" or a codebook matrix
- data
- a data frame or matrix of input data
- sBase
- an object of class "sBase"
- base.color
- short name for the colormap used to encode bases (in row side bar). It can be one of "jet" (jet colormap), "bwr" (blue-white-red colormap), "gbr" (green-black-red colormap), "wyr" (white-yellow-red colormap), "br" (black-red colormap), "yr" (yellow-red colormap), "wb" (white-black colormap), and "rainbow" (rainbow colormap, that is, red-yellow-green-cyan-blue-magenta). Alternatively, any hyphen-separated HTML color names, e.g. "blue-black-yellow", "royalblue-white-sandybrown", "darkgreen-white-darkviolet". A list of standard color names can be found in http://html-color-codes.info/color-names
- base.separated.arg
- a list of main parameters used for styling bar separated lines. See 'Note' below for details on the parameters
- base.legend.location
- location of legend to describe bases. If "none", this legend will not be displayed
- reorderRow
- the way to reorder the rows within a base. It can be "none" for rows within a base being reorded by the hexagon indexes, "hclust" for rows within a base being reorded according to hierarchical clustering of patterns seen, "svd" for rows within a base being reorded according to svd of patterns seen
- keep.data
- logical to indicate whether or not to also write out the input data. By default, it sets to false for not keeping it. It is highly expensive to keep the large data sets
- ...
- additional graphic parameters used in "visHeatmapAdv". For most parameters, please refer to https://www.rdocumentation.org/packages/gplots/topics/heatmap.2
Value
a data frame with following components:
ID: ID for data. It inherits the rownames of data (if exists). Otherwise, it is sequential integer values starting with 1 and ending with dlen, the total number of rows of the input dataHexagon_index: the index for best-matching hexagonsCluster_base: optional, it is only appended when sBase is given. It stores the cluster memberships/basesdata: optional, it is only appended when keep.data is true
Note: the returned data has rows in the same order as visualised in the heatmap
Note
A list of parameters in "base.separated.arg":
- "lty": the line type. Line types can either be specified as an integer (0=blank, 1=solid (default), 2=dashed, 3=dotted, 4=dotdash, 5=longdash, 6=twodash) or as one of the character strings "blank","solid","dashed","dotted","dotdash","longdash","twodash", where "blank" uses 'invisible lines' (i.e., does not draw them)
- "lwd": the line width
- "col": the line color
Examples
# 1) generate an iid normal random matrix of 100x10 data <- matrix( rnorm(100*10,mean=0,sd=1), nrow=100, ncol=10) # 2) get trained using by default setup sMap <- sPipeline(data=data)Start at 2018-01-18 16:56:17 First, define topology of a map grid (2018-01-18 16:56:17)... Second, initialise the codebook matrix (61 X 10) using 'linear' initialisation, given a topology and input data (2018-01-18 16:56:17)... Third, get training at the rough stage (2018-01-18 16:56:17)... 1 out of 7 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 2 out of 7 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 3 out of 7 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 4 out of 7 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 5 out of 7 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 6 out of 7 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 7 out of 7 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) Fourth, get training at the finetune stage (2018-01-18 16:56:17)... 1 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 2 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 3 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 4 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 5 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 6 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 7 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 8 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 9 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 10 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 11 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 12 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 13 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 14 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 15 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 16 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 17 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 18 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 19 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 20 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 21 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 22 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 23 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 24 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) 25 out of 25 (2018-01-18 16:56:17) updated (2018-01-18 16:56:17) Next, identify the best-matching hexagon/rectangle for the input data (2018-01-18 16:56:17)... Finally, append the response data (hits and mqe) into the sMap object (2018-01-18 16:56:17)... Below are the summaries of the training results: dimension of input data: 100x10 xy-dimension of map grid: xdim=9, ydim=9, r=5 grid lattice: hexa grid shape: suprahex dimension of grid coord: 61x2 initialisation method: linear dimension of codebook matrix: 61x10 mean quantization error: 5.02930130271987 Below are the details of trainology: training algorithm: batch alpha type: invert training neighborhood kernel: gaussian trainlength (x input data length): 7 at rough stage; 25 at finetune stage radius (at rough stage): from 3 to 1 radius (at finetune stage): from 1 to 1 End at 2018-01-18 16:56:17 Runtime in total is: 0 secs# 3) partition the grid map into clusters using region-growing algorithm sBase <- sDmatCluster(sMap=sMap, which_neigh=1, distMeasure="median", clusterLinkage="average") # 4) heatmap visualisation output <- visDmatHeatmap(sMap, data, sBase, base.legend.location="bottomleft", labRow=NA)
Source code
visDmatHeatmap.r
Source man
visDmatHeatmap.Rd
visDmatHeatmap.pdf
See also
sDmatCluster, visHeatmapAdv