 Start at 2017-03-27 18:59:46
First, define topology of a map grid (2017-03-27 18:59:46)...
Second, initialise the codebook matrix (169 X 6) using 'uniform' initialisation, given a topology and input data (2017-03-27 18:59:46)...
Third, get training at the rough stage (2017-03-27 18:59:46)...
1 out of 2 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
2 out of 2 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
Fourth, get training at the finetune stage (2017-03-27 18:59:46)...
1 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
2 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
3 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
4 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
5 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
6 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
7 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
Next, identify the best-matching hexagon/rectangle for the input data (2017-03-27 18:59:46)...
Finally, append the response data (hits and mqe) into the sMap object (2017-03-27 18:59:46)...
Below are the summaries of the training results:
dimension of input data: 1000x6
xy-dimension of map grid: xdim=15, ydim=15, r=8
grid lattice: hexa
grid shape: suprahex
dimension of grid coord: 169x2
initialisation method: uniform
dimension of codebook matrix: 169x6
mean quantization error: 1.95201775253281
Below are the details of trainology:
training algorithm: batch
alpha type: invert
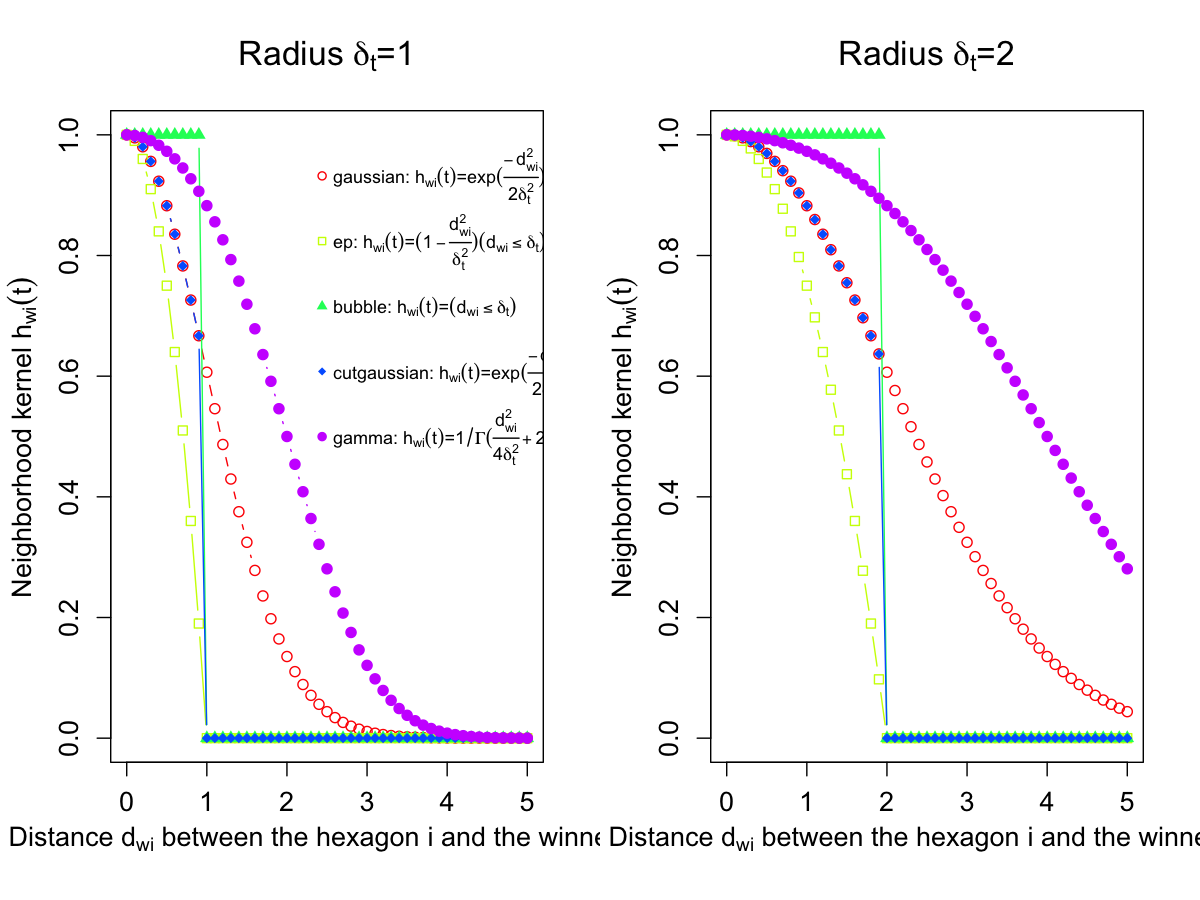
training neighborhood kernel: gaussian
trainlength (x input data length): 2 at rough stage; 7 at finetune stage
radius (at rough stage): from 4 to 1
radius (at finetune stage): from 1 to 1
End at 2017-03-27 18:59:46
Runtime in total is: 0 secs
Start at 2017-03-27 18:59:47
First, define topology of a map grid (2017-03-27 18:59:47)...
Second, initialise the codebook matrix (169 X 6) using 'uniform' initialisation, given a topology and input data (2017-03-27 18:59:47)...
Third, get training at the rough stage (2017-03-27 18:59:47)...
1 out of 2 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
2 out of 2 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
Fourth, get training at the finetune stage (2017-03-27 18:59:47)...
1 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
2 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
3 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
4 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
5 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
6 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
7 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
Next, identify the best-matching hexagon/rectangle for the input data (2017-03-27 18:59:47)...
Finally, append the response data (hits and mqe) into the sMap object (2017-03-27 18:59:47)...
Below are the summaries of the training results:
dimension of input data: 1000x6
xy-dimension of map grid: xdim=15, ydim=15, r=8
grid lattice: hexa
grid shape: suprahex
dimension of grid coord: 169x2
initialisation method: uniform
dimension of codebook matrix: 169x6
mean quantization error: 2.3688468448732
Below are the details of trainology:
training algorithm: batch
alpha type: invert
training neighborhood kernel: gamma
trainlength (x input data length): 2 at rough stage; 7 at finetune stage
radius (at rough stage): from 4 to 1
radius (at finetune stage): from 1 to 1
End at 2017-03-27 18:59:47
Runtime in total is: 0 secs
Start at 2017-03-27 18:59:46
First, define topology of a map grid (2017-03-27 18:59:46)...
Second, initialise the codebook matrix (169 X 6) using 'uniform' initialisation, given a topology and input data (2017-03-27 18:59:46)...
Third, get training at the rough stage (2017-03-27 18:59:46)...
1 out of 2 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
2 out of 2 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
Fourth, get training at the finetune stage (2017-03-27 18:59:46)...
1 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
2 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
3 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
4 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
5 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
6 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
7 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
Next, identify the best-matching hexagon/rectangle for the input data (2017-03-27 18:59:46)...
Finally, append the response data (hits and mqe) into the sMap object (2017-03-27 18:59:46)...
Below are the summaries of the training results:
dimension of input data: 1000x6
xy-dimension of map grid: xdim=15, ydim=15, r=8
grid lattice: hexa
grid shape: suprahex
dimension of grid coord: 169x2
initialisation method: uniform
dimension of codebook matrix: 169x6
mean quantization error: 1.95201775253281
Below are the details of trainology:
training algorithm: batch
alpha type: invert
training neighborhood kernel: gaussian
trainlength (x input data length): 2 at rough stage; 7 at finetune stage
radius (at rough stage): from 4 to 1
radius (at finetune stage): from 1 to 1
End at 2017-03-27 18:59:46
Runtime in total is: 0 secs
Start at 2017-03-27 18:59:47
First, define topology of a map grid (2017-03-27 18:59:47)...
Second, initialise the codebook matrix (169 X 6) using 'uniform' initialisation, given a topology and input data (2017-03-27 18:59:47)...
Third, get training at the rough stage (2017-03-27 18:59:47)...
1 out of 2 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
2 out of 2 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
Fourth, get training at the finetune stage (2017-03-27 18:59:47)...
1 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
2 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
3 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
4 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
5 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
6 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
7 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
Next, identify the best-matching hexagon/rectangle for the input data (2017-03-27 18:59:47)...
Finally, append the response data (hits and mqe) into the sMap object (2017-03-27 18:59:47)...
Below are the summaries of the training results:
dimension of input data: 1000x6
xy-dimension of map grid: xdim=15, ydim=15, r=8
grid lattice: hexa
grid shape: suprahex
dimension of grid coord: 169x2
initialisation method: uniform
dimension of codebook matrix: 169x6
mean quantization error: 2.3688468448732
Below are the details of trainology:
training algorithm: batch
alpha type: invert
training neighborhood kernel: gamma
trainlength (x input data length): 2 at rough stage; 7 at finetune stage
radius (at rough stage): from 4 to 1
radius (at finetune stage): from 1 to 1
End at 2017-03-27 18:59:47
Runtime in total is: 0 secs
 Start at 2017-03-27 18:59:48
First, define topology of a map grid (2017-03-27 18:59:48)...
Second, initialise the codebook matrix (169 X 6) using 'uniform' initialisation, given a topology and input data (2017-03-27 18:59:48)...
Third, get training at the rough stage (2017-03-27 18:59:48)...
1 out of 2 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
2 out of 2 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
Fourth, get training at the finetune stage (2017-03-27 18:59:48)...
1 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
2 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
3 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
4 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
5 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
6 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
7 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
Next, identify the best-matching hexagon/rectangle for the input data (2017-03-27 18:59:48)...
Finally, append the response data (hits and mqe) into the sMap object (2017-03-27 18:59:48)...
Below are the summaries of the training results:
dimension of input data: 1000x6
xy-dimension of map grid: xdim=15, ydim=15, r=8
grid lattice: hexa
grid shape: suprahex
dimension of grid coord: 169x2
initialisation method: uniform
dimension of codebook matrix: 169x6
mean quantization error: 0.97279930292498
Below are the details of trainology:
training algorithm: batch
alpha type: invert
training neighborhood kernel: ep
trainlength (x input data length): 2 at rough stage; 7 at finetune stage
radius (at rough stage): from 4 to 1
radius (at finetune stage): from 1 to 1
End at 2017-03-27 18:59:48
Runtime in total is: 0 secs
Start at 2017-03-27 18:59:48
First, define topology of a map grid (2017-03-27 18:59:48)...
Second, initialise the codebook matrix (169 X 6) using 'uniform' initialisation, given a topology and input data (2017-03-27 18:59:48)...
Third, get training at the rough stage (2017-03-27 18:59:48)...
1 out of 2 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
2 out of 2 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
Fourth, get training at the finetune stage (2017-03-27 18:59:48)...
1 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
2 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
3 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
4 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
5 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
6 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
7 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
Next, identify the best-matching hexagon/rectangle for the input data (2017-03-27 18:59:48)...
Finally, append the response data (hits and mqe) into the sMap object (2017-03-27 18:59:48)...
Below are the summaries of the training results:
dimension of input data: 1000x6
xy-dimension of map grid: xdim=15, ydim=15, r=8
grid lattice: hexa
grid shape: suprahex
dimension of grid coord: 169x2
initialisation method: uniform
dimension of codebook matrix: 169x6
mean quantization error: 0.97279930292498
Below are the details of trainology:
training algorithm: batch
alpha type: invert
training neighborhood kernel: ep
trainlength (x input data length): 2 at rough stage; 7 at finetune stage
radius (at rough stage): from 4 to 1
radius (at finetune stage): from 1 to 1
End at 2017-03-27 18:59:48
Runtime in total is: 0 secs
 Start at 2017-03-27 18:59:49
First, define topology of a map grid (2017-03-27 18:59:49)...
Second, initialise the codebook matrix (169 X 6) using 'uniform' initialisation, given a topology and input data (2017-03-27 18:59:49)...
Third, get training at the rough stage (2017-03-27 18:59:49)...
1 out of 2 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
2 out of 2 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
Fourth, get training at the finetune stage (2017-03-27 18:59:49)...
1 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
2 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
3 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
4 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
5 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
6 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
7 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
Next, identify the best-matching hexagon/rectangle for the input data (2017-03-27 18:59:49)...
Finally, append the response data (hits and mqe) into the sMap object (2017-03-27 18:59:49)...
Below are the summaries of the training results:
dimension of input data: 1000x6
xy-dimension of map grid: xdim=15, ydim=15, r=8
grid lattice: hexa
grid shape: suprahex
dimension of grid coord: 169x2
initialisation method: uniform
dimension of codebook matrix: 169x6
mean quantization error: 1.52361772550777
Below are the details of trainology:
training algorithm: batch
alpha type: invert
training neighborhood kernel: cutgaussian
trainlength (x input data length): 2 at rough stage; 7 at finetune stage
radius (at rough stage): from 4 to 1
radius (at finetune stage): from 1 to 1
End at 2017-03-27 18:59:49
Runtime in total is: 0 secs
Start at 2017-03-27 18:59:49
First, define topology of a map grid (2017-03-27 18:59:49)...
Second, initialise the codebook matrix (169 X 6) using 'uniform' initialisation, given a topology and input data (2017-03-27 18:59:49)...
Third, get training at the rough stage (2017-03-27 18:59:49)...
1 out of 2 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
2 out of 2 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
Fourth, get training at the finetune stage (2017-03-27 18:59:49)...
1 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
2 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
3 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
4 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
5 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
6 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
7 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
Next, identify the best-matching hexagon/rectangle for the input data (2017-03-27 18:59:49)...
Finally, append the response data (hits and mqe) into the sMap object (2017-03-27 18:59:49)...
Below are the summaries of the training results:
dimension of input data: 1000x6
xy-dimension of map grid: xdim=15, ydim=15, r=8
grid lattice: hexa
grid shape: suprahex
dimension of grid coord: 169x2
initialisation method: uniform
dimension of codebook matrix: 169x6
mean quantization error: 1.52361772550777
Below are the details of trainology:
training algorithm: batch
alpha type: invert
training neighborhood kernel: cutgaussian
trainlength (x input data length): 2 at rough stage; 7 at finetune stage
radius (at rough stage): from 4 to 1
radius (at finetune stage): from 1 to 1
End at 2017-03-27 18:59:49
Runtime in total is: 0 secs
 Start at 2017-03-27 18:59:49
First, define topology of a map grid (2017-03-27 18:59:49)...
Second, initialise the codebook matrix (169 X 6) using 'uniform' initialisation, given a topology and input data (2017-03-27 18:59:49)...
Third, get training at the rough stage (2017-03-27 18:59:49)...
1 out of 2 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
2 out of 2 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:50)
Fourth, get training at the finetune stage (2017-03-27 18:59:50)...
1 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
2 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
3 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
4 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
5 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
6 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
7 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
Next, identify the best-matching hexagon/rectangle for the input data (2017-03-27 18:59:50)...
Finally, append the response data (hits and mqe) into the sMap object (2017-03-27 18:59:50)...
Below are the summaries of the training results:
dimension of input data: 1000x6
xy-dimension of map grid: xdim=15, ydim=15, r=8
grid lattice: hexa
grid shape: suprahex
dimension of grid coord: 169x2
initialisation method: uniform
dimension of codebook matrix: 169x6
mean quantization error: 1.59843633585734
Below are the details of trainology:
training algorithm: batch
alpha type: invert
training neighborhood kernel: bubble
trainlength (x input data length): 2 at rough stage; 7 at finetune stage
radius (at rough stage): from 4 to 1
radius (at finetune stage): from 1 to 1
End at 2017-03-27 18:59:50
Runtime in total is: 1 secs
Start at 2017-03-27 18:59:49
First, define topology of a map grid (2017-03-27 18:59:49)...
Second, initialise the codebook matrix (169 X 6) using 'uniform' initialisation, given a topology and input data (2017-03-27 18:59:49)...
Third, get training at the rough stage (2017-03-27 18:59:49)...
1 out of 2 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
2 out of 2 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:50)
Fourth, get training at the finetune stage (2017-03-27 18:59:50)...
1 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
2 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
3 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
4 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
5 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
6 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
7 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
Next, identify the best-matching hexagon/rectangle for the input data (2017-03-27 18:59:50)...
Finally, append the response data (hits and mqe) into the sMap object (2017-03-27 18:59:50)...
Below are the summaries of the training results:
dimension of input data: 1000x6
xy-dimension of map grid: xdim=15, ydim=15, r=8
grid lattice: hexa
grid shape: suprahex
dimension of grid coord: 169x2
initialisation method: uniform
dimension of codebook matrix: 169x6
mean quantization error: 1.59843633585734
Below are the details of trainology:
training algorithm: batch
alpha type: invert
training neighborhood kernel: bubble
trainlength (x input data length): 2 at rough stage; 7 at finetune stage
radius (at rough stage): from 4 to 1
radius (at finetune stage): from 1 to 1
End at 2017-03-27 18:59:50
Runtime in total is: 1 secs
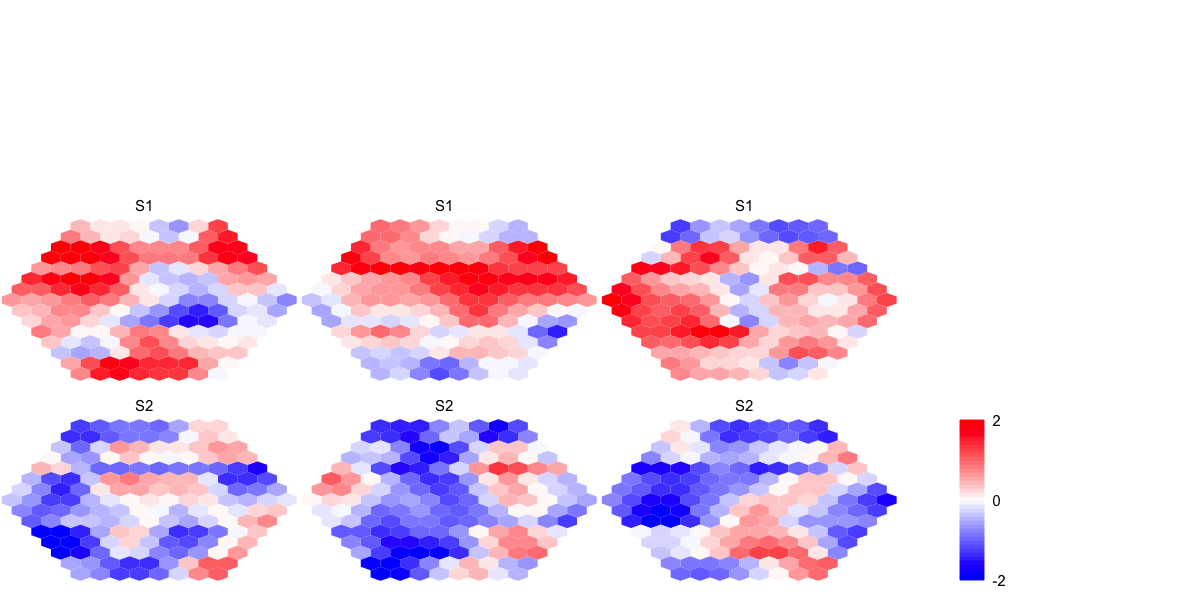
 colnames(data) <- c("S1","S1","S1","S2","S2","S2")
# with "gaussian" kernel (by default)
sMap_ga <- sPipeline(data=data, neighKernel="gaussian", init="uniform")
Start at 2017-03-27 18:59:46
First, define topology of a map grid (2017-03-27 18:59:46)...
Second, initialise the codebook matrix (169 X 6) using 'uniform' initialisation, given a topology and input data (2017-03-27 18:59:46)...
Third, get training at the rough stage (2017-03-27 18:59:46)...
1 out of 2 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
2 out of 2 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
Fourth, get training at the finetune stage (2017-03-27 18:59:46)...
1 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
2 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
3 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
4 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
5 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
6 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
7 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
Next, identify the best-matching hexagon/rectangle for the input data (2017-03-27 18:59:46)...
Finally, append the response data (hits and mqe) into the sMap object (2017-03-27 18:59:46)...
Below are the summaries of the training results:
dimension of input data: 1000x6
xy-dimension of map grid: xdim=15, ydim=15, r=8
grid lattice: hexa
grid shape: suprahex
dimension of grid coord: 169x2
initialisation method: uniform
dimension of codebook matrix: 169x6
mean quantization error: 1.95201775253281
Below are the details of trainology:
training algorithm: batch
alpha type: invert
training neighborhood kernel: gaussian
trainlength (x input data length): 2 at rough stage; 7 at finetune stage
radius (at rough stage): from 4 to 1
radius (at finetune stage): from 1 to 1
End at 2017-03-27 18:59:46
Runtime in total is: 0 secs
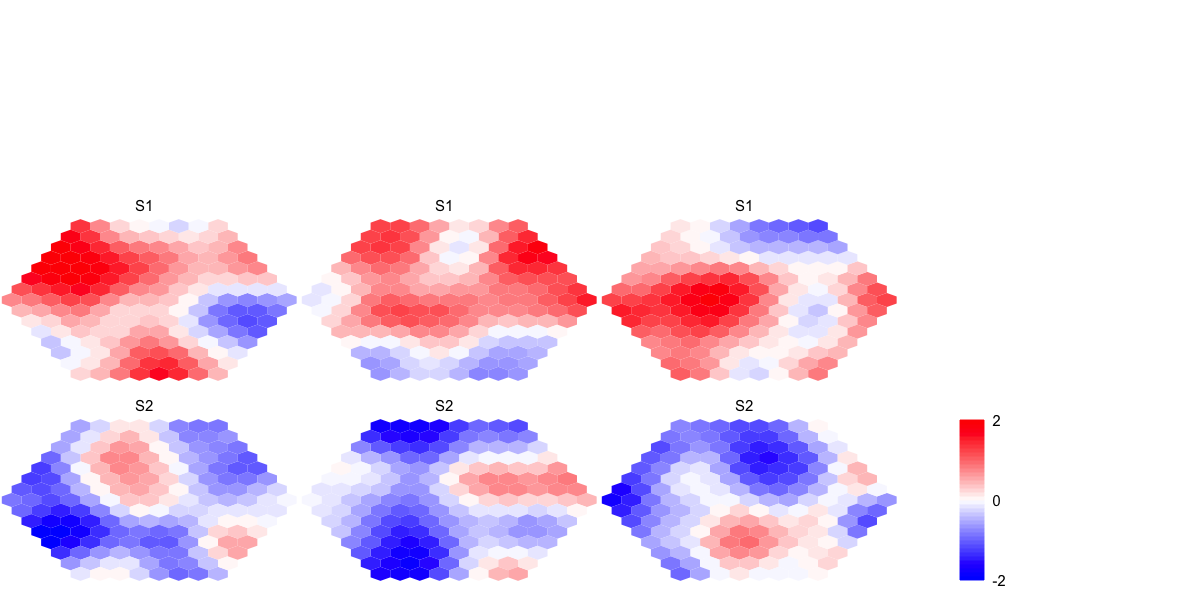
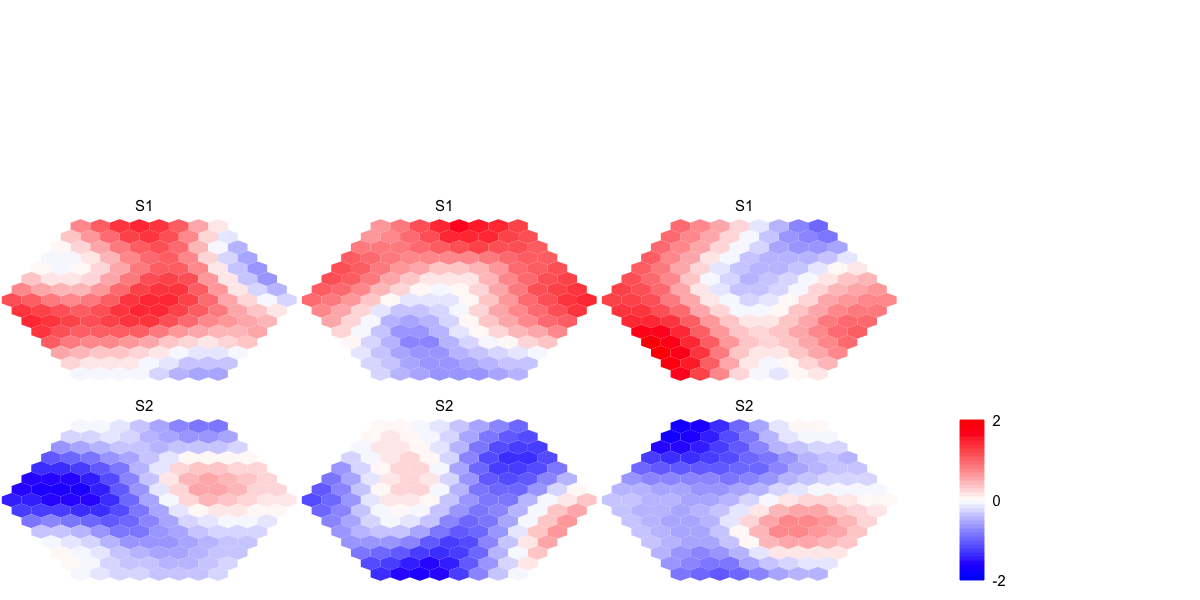
visHexMulComp(sMap_ga)
# with "gamma" kernel
sMap_gm <- sPipeline(data=data, neighKernel="gamma", init="uniform")
Start at 2017-03-27 18:59:47
First, define topology of a map grid (2017-03-27 18:59:47)...
Second, initialise the codebook matrix (169 X 6) using 'uniform' initialisation, given a topology and input data (2017-03-27 18:59:47)...
Third, get training at the rough stage (2017-03-27 18:59:47)...
1 out of 2 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
2 out of 2 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
Fourth, get training at the finetune stage (2017-03-27 18:59:47)...
1 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
2 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
3 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
4 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
5 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
6 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
7 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
Next, identify the best-matching hexagon/rectangle for the input data (2017-03-27 18:59:47)...
Finally, append the response data (hits and mqe) into the sMap object (2017-03-27 18:59:47)...
Below are the summaries of the training results:
dimension of input data: 1000x6
xy-dimension of map grid: xdim=15, ydim=15, r=8
grid lattice: hexa
grid shape: suprahex
dimension of grid coord: 169x2
initialisation method: uniform
dimension of codebook matrix: 169x6
mean quantization error: 2.3688468448732
Below are the details of trainology:
training algorithm: batch
alpha type: invert
training neighborhood kernel: gamma
trainlength (x input data length): 2 at rough stage; 7 at finetune stage
radius (at rough stage): from 4 to 1
radius (at finetune stage): from 1 to 1
End at 2017-03-27 18:59:47
Runtime in total is: 0 secs
colnames(data) <- c("S1","S1","S1","S2","S2","S2")
# with "gaussian" kernel (by default)
sMap_ga <- sPipeline(data=data, neighKernel="gaussian", init="uniform")
Start at 2017-03-27 18:59:46
First, define topology of a map grid (2017-03-27 18:59:46)...
Second, initialise the codebook matrix (169 X 6) using 'uniform' initialisation, given a topology and input data (2017-03-27 18:59:46)...
Third, get training at the rough stage (2017-03-27 18:59:46)...
1 out of 2 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
2 out of 2 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
Fourth, get training at the finetune stage (2017-03-27 18:59:46)...
1 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
2 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
3 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
4 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
5 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
6 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
7 out of 7 (2017-03-27 18:59:46)
updated (2017-03-27 18:59:46)
Next, identify the best-matching hexagon/rectangle for the input data (2017-03-27 18:59:46)...
Finally, append the response data (hits and mqe) into the sMap object (2017-03-27 18:59:46)...
Below are the summaries of the training results:
dimension of input data: 1000x6
xy-dimension of map grid: xdim=15, ydim=15, r=8
grid lattice: hexa
grid shape: suprahex
dimension of grid coord: 169x2
initialisation method: uniform
dimension of codebook matrix: 169x6
mean quantization error: 1.95201775253281
Below are the details of trainology:
training algorithm: batch
alpha type: invert
training neighborhood kernel: gaussian
trainlength (x input data length): 2 at rough stage; 7 at finetune stage
radius (at rough stage): from 4 to 1
radius (at finetune stage): from 1 to 1
End at 2017-03-27 18:59:46
Runtime in total is: 0 secs
visHexMulComp(sMap_ga)
# with "gamma" kernel
sMap_gm <- sPipeline(data=data, neighKernel="gamma", init="uniform")
Start at 2017-03-27 18:59:47
First, define topology of a map grid (2017-03-27 18:59:47)...
Second, initialise the codebook matrix (169 X 6) using 'uniform' initialisation, given a topology and input data (2017-03-27 18:59:47)...
Third, get training at the rough stage (2017-03-27 18:59:47)...
1 out of 2 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
2 out of 2 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
Fourth, get training at the finetune stage (2017-03-27 18:59:47)...
1 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
2 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
3 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
4 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
5 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
6 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
7 out of 7 (2017-03-27 18:59:47)
updated (2017-03-27 18:59:47)
Next, identify the best-matching hexagon/rectangle for the input data (2017-03-27 18:59:47)...
Finally, append the response data (hits and mqe) into the sMap object (2017-03-27 18:59:47)...
Below are the summaries of the training results:
dimension of input data: 1000x6
xy-dimension of map grid: xdim=15, ydim=15, r=8
grid lattice: hexa
grid shape: suprahex
dimension of grid coord: 169x2
initialisation method: uniform
dimension of codebook matrix: 169x6
mean quantization error: 2.3688468448732
Below are the details of trainology:
training algorithm: batch
alpha type: invert
training neighborhood kernel: gamma
trainlength (x input data length): 2 at rough stage; 7 at finetune stage
radius (at rough stage): from 4 to 1
radius (at finetune stage): from 1 to 1
End at 2017-03-27 18:59:47
Runtime in total is: 0 secs
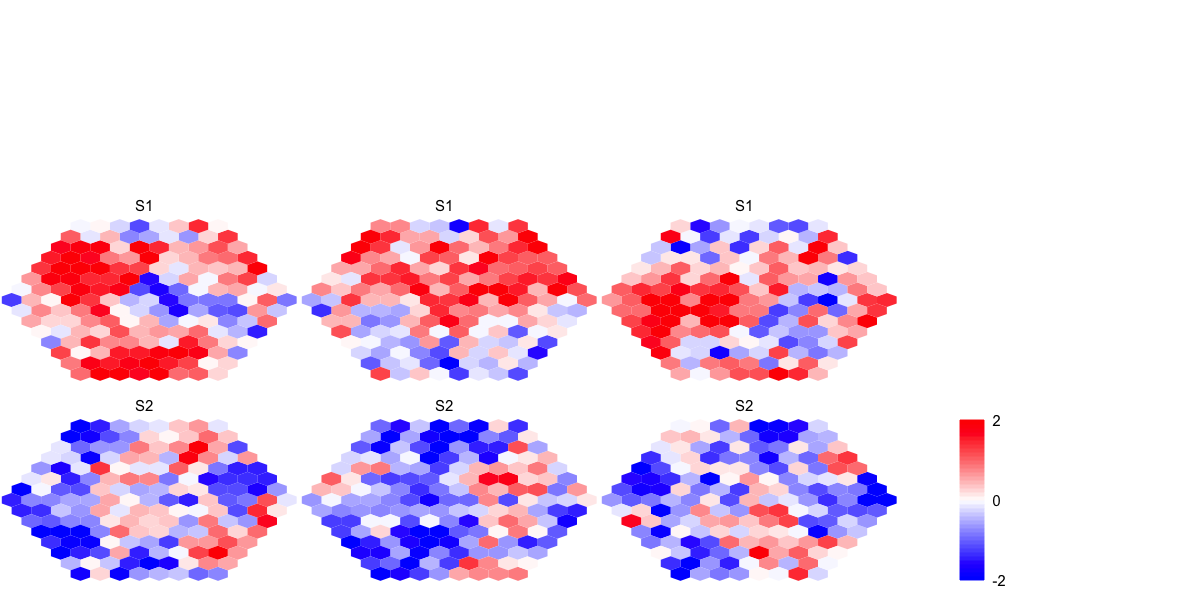
 visHexMulComp(sMap_gm)
# with "ep" kernel
sMap_ep <- sPipeline(data=data, neighKernel="ep", init="uniform")
Start at 2017-03-27 18:59:48
First, define topology of a map grid (2017-03-27 18:59:48)...
Second, initialise the codebook matrix (169 X 6) using 'uniform' initialisation, given a topology and input data (2017-03-27 18:59:48)...
Third, get training at the rough stage (2017-03-27 18:59:48)...
1 out of 2 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
2 out of 2 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
Fourth, get training at the finetune stage (2017-03-27 18:59:48)...
1 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
2 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
3 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
4 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
5 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
6 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
7 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
Next, identify the best-matching hexagon/rectangle for the input data (2017-03-27 18:59:48)...
Finally, append the response data (hits and mqe) into the sMap object (2017-03-27 18:59:48)...
Below are the summaries of the training results:
dimension of input data: 1000x6
xy-dimension of map grid: xdim=15, ydim=15, r=8
grid lattice: hexa
grid shape: suprahex
dimension of grid coord: 169x2
initialisation method: uniform
dimension of codebook matrix: 169x6
mean quantization error: 0.97279930292498
Below are the details of trainology:
training algorithm: batch
alpha type: invert
training neighborhood kernel: ep
trainlength (x input data length): 2 at rough stage; 7 at finetune stage
radius (at rough stage): from 4 to 1
radius (at finetune stage): from 1 to 1
End at 2017-03-27 18:59:48
Runtime in total is: 0 secs
visHexMulComp(sMap_gm)
# with "ep" kernel
sMap_ep <- sPipeline(data=data, neighKernel="ep", init="uniform")
Start at 2017-03-27 18:59:48
First, define topology of a map grid (2017-03-27 18:59:48)...
Second, initialise the codebook matrix (169 X 6) using 'uniform' initialisation, given a topology and input data (2017-03-27 18:59:48)...
Third, get training at the rough stage (2017-03-27 18:59:48)...
1 out of 2 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
2 out of 2 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
Fourth, get training at the finetune stage (2017-03-27 18:59:48)...
1 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
2 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
3 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
4 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
5 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
6 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
7 out of 7 (2017-03-27 18:59:48)
updated (2017-03-27 18:59:48)
Next, identify the best-matching hexagon/rectangle for the input data (2017-03-27 18:59:48)...
Finally, append the response data (hits and mqe) into the sMap object (2017-03-27 18:59:48)...
Below are the summaries of the training results:
dimension of input data: 1000x6
xy-dimension of map grid: xdim=15, ydim=15, r=8
grid lattice: hexa
grid shape: suprahex
dimension of grid coord: 169x2
initialisation method: uniform
dimension of codebook matrix: 169x6
mean quantization error: 0.97279930292498
Below are the details of trainology:
training algorithm: batch
alpha type: invert
training neighborhood kernel: ep
trainlength (x input data length): 2 at rough stage; 7 at finetune stage
radius (at rough stage): from 4 to 1
radius (at finetune stage): from 1 to 1
End at 2017-03-27 18:59:48
Runtime in total is: 0 secs
 visHexMulComp(sMap_ep)
# with "cutgaussian" kernel
sMap_cu <- sPipeline(data=data, neighKernel="cutgaussian", init="uniform")
Start at 2017-03-27 18:59:49
First, define topology of a map grid (2017-03-27 18:59:49)...
Second, initialise the codebook matrix (169 X 6) using 'uniform' initialisation, given a topology and input data (2017-03-27 18:59:49)...
Third, get training at the rough stage (2017-03-27 18:59:49)...
1 out of 2 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
2 out of 2 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
Fourth, get training at the finetune stage (2017-03-27 18:59:49)...
1 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
2 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
3 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
4 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
5 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
6 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
7 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
Next, identify the best-matching hexagon/rectangle for the input data (2017-03-27 18:59:49)...
Finally, append the response data (hits and mqe) into the sMap object (2017-03-27 18:59:49)...
Below are the summaries of the training results:
dimension of input data: 1000x6
xy-dimension of map grid: xdim=15, ydim=15, r=8
grid lattice: hexa
grid shape: suprahex
dimension of grid coord: 169x2
initialisation method: uniform
dimension of codebook matrix: 169x6
mean quantization error: 1.52361772550777
Below are the details of trainology:
training algorithm: batch
alpha type: invert
training neighborhood kernel: cutgaussian
trainlength (x input data length): 2 at rough stage; 7 at finetune stage
radius (at rough stage): from 4 to 1
radius (at finetune stage): from 1 to 1
End at 2017-03-27 18:59:49
Runtime in total is: 0 secs
visHexMulComp(sMap_ep)
# with "cutgaussian" kernel
sMap_cu <- sPipeline(data=data, neighKernel="cutgaussian", init="uniform")
Start at 2017-03-27 18:59:49
First, define topology of a map grid (2017-03-27 18:59:49)...
Second, initialise the codebook matrix (169 X 6) using 'uniform' initialisation, given a topology and input data (2017-03-27 18:59:49)...
Third, get training at the rough stage (2017-03-27 18:59:49)...
1 out of 2 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
2 out of 2 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
Fourth, get training at the finetune stage (2017-03-27 18:59:49)...
1 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
2 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
3 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
4 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
5 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
6 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
7 out of 7 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
Next, identify the best-matching hexagon/rectangle for the input data (2017-03-27 18:59:49)...
Finally, append the response data (hits and mqe) into the sMap object (2017-03-27 18:59:49)...
Below are the summaries of the training results:
dimension of input data: 1000x6
xy-dimension of map grid: xdim=15, ydim=15, r=8
grid lattice: hexa
grid shape: suprahex
dimension of grid coord: 169x2
initialisation method: uniform
dimension of codebook matrix: 169x6
mean quantization error: 1.52361772550777
Below are the details of trainology:
training algorithm: batch
alpha type: invert
training neighborhood kernel: cutgaussian
trainlength (x input data length): 2 at rough stage; 7 at finetune stage
radius (at rough stage): from 4 to 1
radius (at finetune stage): from 1 to 1
End at 2017-03-27 18:59:49
Runtime in total is: 0 secs
 visHexMulComp(sMap_cu)
# with "bubble" kernel
sMap_bu <- sPipeline(data=data, neighKernel="bubble", init="uniform")
Start at 2017-03-27 18:59:49
First, define topology of a map grid (2017-03-27 18:59:49)...
Second, initialise the codebook matrix (169 X 6) using 'uniform' initialisation, given a topology and input data (2017-03-27 18:59:49)...
Third, get training at the rough stage (2017-03-27 18:59:49)...
1 out of 2 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
2 out of 2 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:50)
Fourth, get training at the finetune stage (2017-03-27 18:59:50)...
1 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
2 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
3 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
4 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
5 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
6 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
7 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
Next, identify the best-matching hexagon/rectangle for the input data (2017-03-27 18:59:50)...
Finally, append the response data (hits and mqe) into the sMap object (2017-03-27 18:59:50)...
Below are the summaries of the training results:
dimension of input data: 1000x6
xy-dimension of map grid: xdim=15, ydim=15, r=8
grid lattice: hexa
grid shape: suprahex
dimension of grid coord: 169x2
initialisation method: uniform
dimension of codebook matrix: 169x6
mean quantization error: 1.59843633585734
Below are the details of trainology:
training algorithm: batch
alpha type: invert
training neighborhood kernel: bubble
trainlength (x input data length): 2 at rough stage; 7 at finetune stage
radius (at rough stage): from 4 to 1
radius (at finetune stage): from 1 to 1
End at 2017-03-27 18:59:50
Runtime in total is: 1 secs
visHexMulComp(sMap_cu)
# with "bubble" kernel
sMap_bu <- sPipeline(data=data, neighKernel="bubble", init="uniform")
Start at 2017-03-27 18:59:49
First, define topology of a map grid (2017-03-27 18:59:49)...
Second, initialise the codebook matrix (169 X 6) using 'uniform' initialisation, given a topology and input data (2017-03-27 18:59:49)...
Third, get training at the rough stage (2017-03-27 18:59:49)...
1 out of 2 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:49)
2 out of 2 (2017-03-27 18:59:49)
updated (2017-03-27 18:59:50)
Fourth, get training at the finetune stage (2017-03-27 18:59:50)...
1 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
2 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
3 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
4 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
5 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
6 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
7 out of 7 (2017-03-27 18:59:50)
updated (2017-03-27 18:59:50)
Next, identify the best-matching hexagon/rectangle for the input data (2017-03-27 18:59:50)...
Finally, append the response data (hits and mqe) into the sMap object (2017-03-27 18:59:50)...
Below are the summaries of the training results:
dimension of input data: 1000x6
xy-dimension of map grid: xdim=15, ydim=15, r=8
grid lattice: hexa
grid shape: suprahex
dimension of grid coord: 169x2
initialisation method: uniform
dimension of codebook matrix: 169x6
mean quantization error: 1.59843633585734
Below are the details of trainology:
training algorithm: batch
alpha type: invert
training neighborhood kernel: bubble
trainlength (x input data length): 2 at rough stage; 7 at finetune stage
radius (at rough stage): from 4 to 1
radius (at finetune stage): from 1 to 1
End at 2017-03-27 18:59:50
Runtime in total is: 1 secs
 visHexMulComp(sMap_bu)
visHexMulComp(sMap_bu)